Research
Multimodal Biomedical data analysis
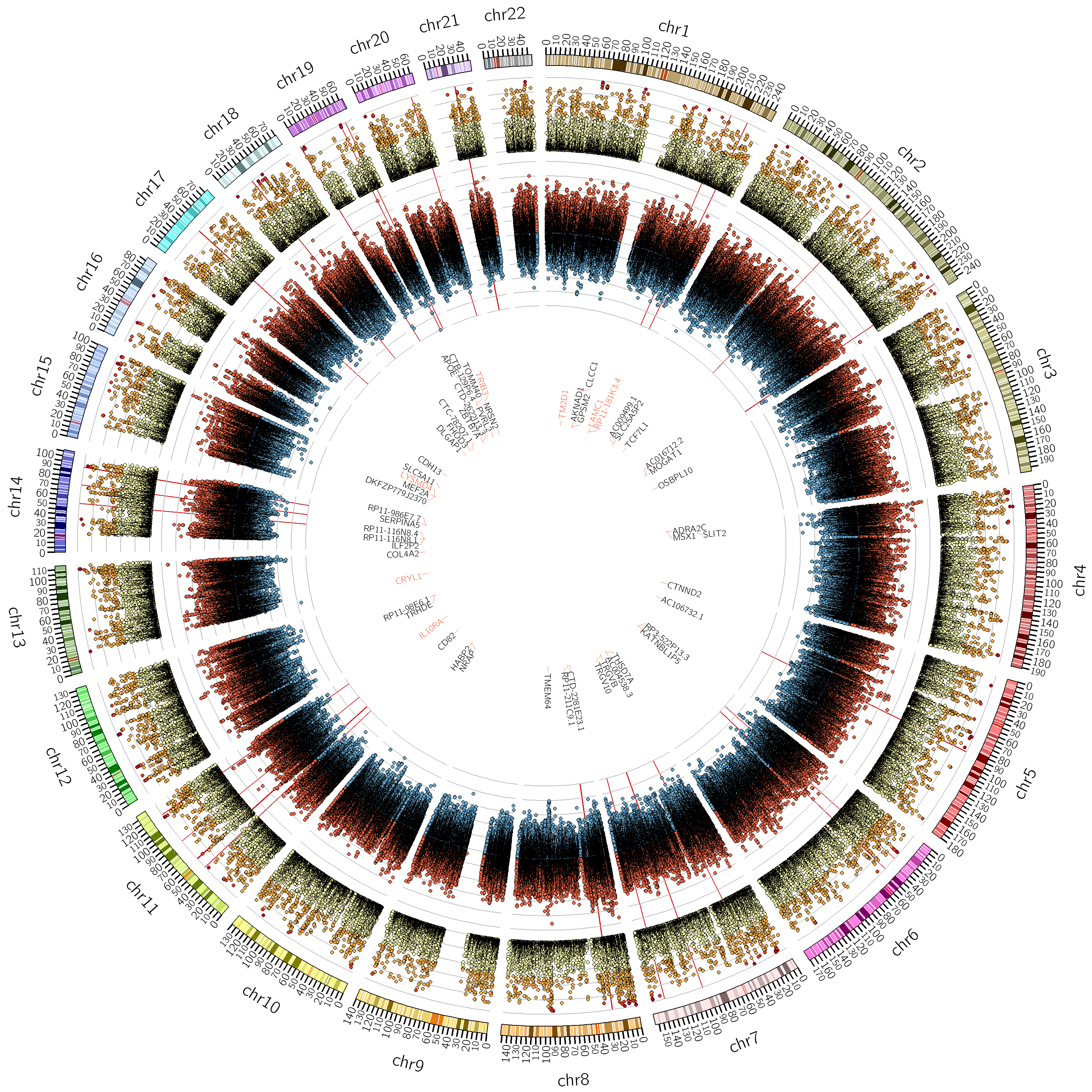
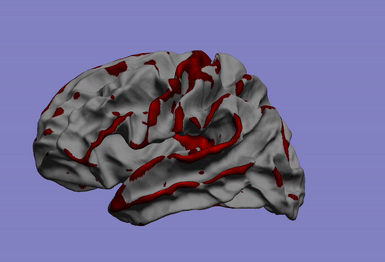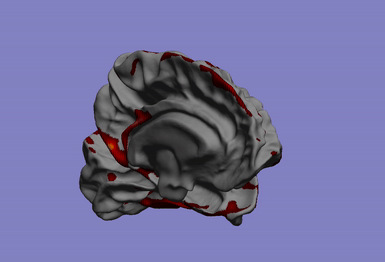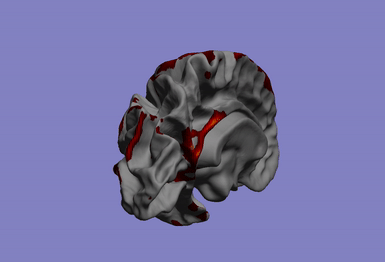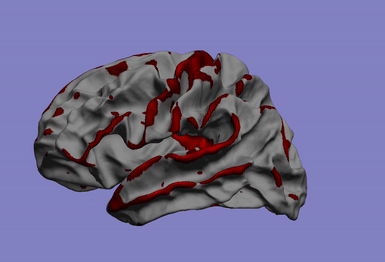
This research axis focuses on the study of the relationship between complex biomedical data, such as phenotypes quantified by neuroimaging (such as atrophy, or connectivity) and individual’s -omics profile (e.g. large genetic arrays). To solve this problem, we develop efficient and flexible statistical models for studying multivariate patterns of associations between mutimodal biomedical data.
Selected software and publications:
Spatio-temporal analysis of biomedical data
I study learning algorithms for modeling and predicting longitudinal changes in imaging, clinical and biological data.
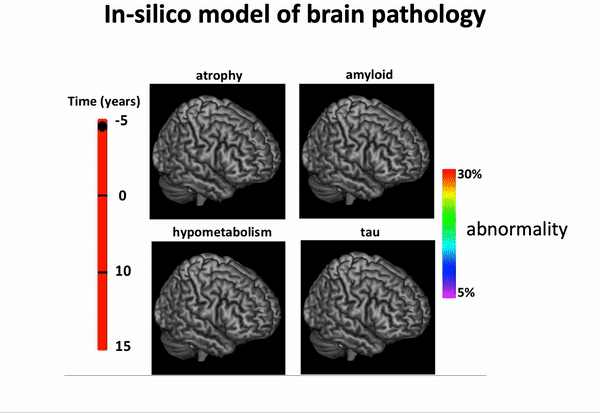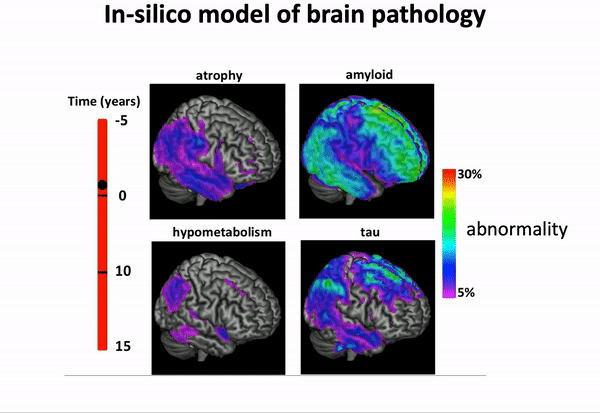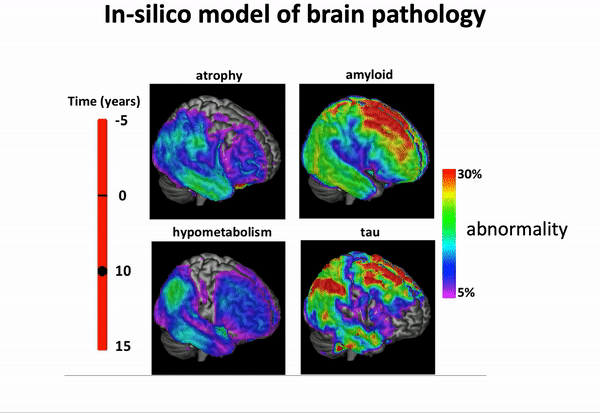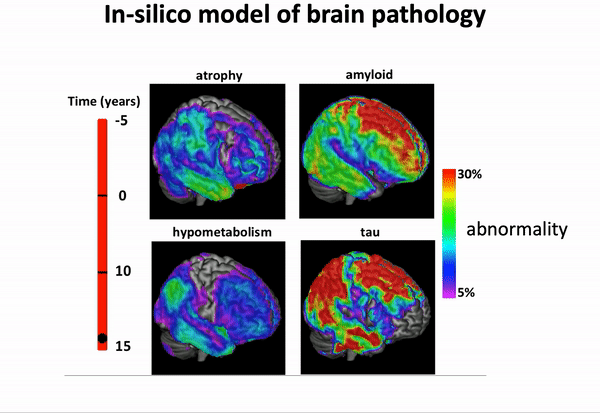
We can think of time series of patients’ data as measurements from an underlying disease progression trajectory spanning several years.
Yet, when we observe the patients during a clinical trial, the disease time information is unknown. This fascinating research topic aims at reconstructing the natural history of a pathology from individual time series of clinical and medical imaging data.
Tutorials:
Software and papers:
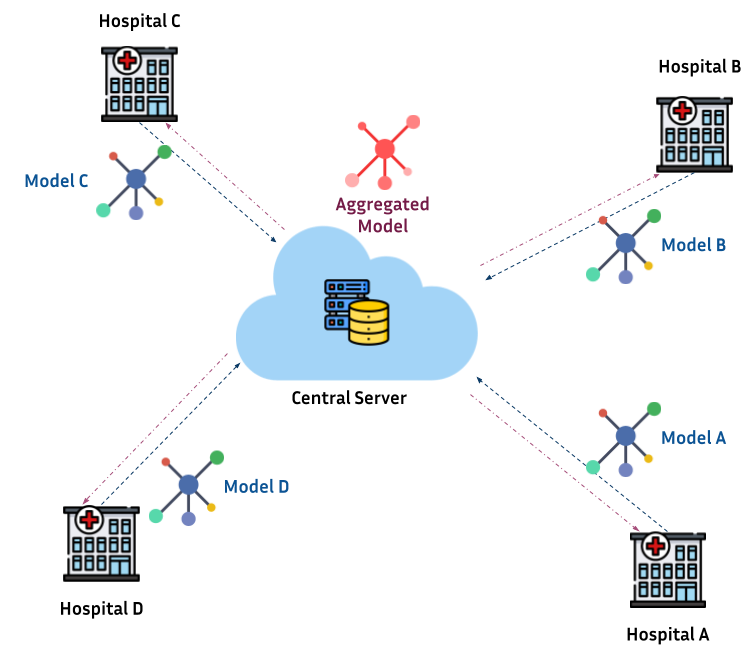
Federated learning in biomedical applications
My research is on the theory and practical use-cases of federated learning (FL), with a special focus in healthcare applications. Current topics include:
- Studying the statistical properties of aggregation mechanisms and clients’ sampling methods in heterogeneity FL setting
- Investigating robust FL methods with respect to clients’ heterogeneity and attacks
- Developing the FL software Fed-BioMed an open source project focused on empowering biomedical research using non-centralized approaches for statistical analysis and machine learning